 Author
Author  Correspondence author
Correspondence author
Molecular Plant Breeding, 2013, Vol. 4, No. 2 doi: 10.5376/mpb.2013.04.0002
Received: 21 Nov., 2012 Accepted: 07 Dec., 2012 Published: 24 Dec., 2012
Liu J.X., et al., 2012, Molecular Cloning and Characterization of the GoMADS-box2 Gene from Guzmania Ruiz & Pav, Molecular Plant Breeding, Vol.4, No.2 15-23 (doi: 10.5376/mpb.2013.04.0002)
The MADS-box gene plays a crucial role in the formation of the floral meristem, development of floral organs and florescence regulation in plants. Based on a full-length cDNA library of floral organs and bracts in ostara, a Guzmania cultivar, four EST sequences from the same contig that included the MADS-box gene were obtained by large-scale random sequencing. Using the primer walking sequencing method, a 985 bp full-length cDNA sequence of the MADS-box gene was identified, which was named GoMADS-box2 gene (GenBank accession No.:JN936044). It has a 675 bp open reading frame that encodes 225 amino acid residuals, and its theoretical molecular weight and isoelectric point were predicted to be 26.3 kD and 9.36, respectively, by using a ProtParam analysis. The putative GoMADS-box2 protein contains a MIKC domain, which only exists in plant MADS-box proteins. The SPOMA program predicted that the secondary structure of the protein was composed of alpha helices (56.00%), random coils (24.44%), extended strands (13.78%) and beta turns (5.78%). Based on the 3D structure of the 1TQE P chain, a tertiary structure model was predicted by the ESyPred 3D program. Molecular phylogenetic tree analysis revealed that the cloned gene belongs to the AP3-type family of genes as well as B-class genes, which might control the development of the petals and stamens. Finally, real-time quantitative PCR analysis indicated that the GoMADS-box2 gene could be expressed in all of the tested tissues, and the amount expressed in the flower was much higher than that of other three tissues; while the expression levels of the scape and leaf were nearly equal and the expression level in the bract was the lowest that was close to zero.
The development of the flower in plants is controlled by a series of genes that are influenced and regulated temporally and spatially by the internal and external factors. The classic ABC model of the flower development and regulation was proposed in 1991 (Coen and Meyerowitz, 1991; Weigel and Meyerowitz, 1994; Pelaz et al., 2000; Honma and Goto, 2001; Theissen and Saedler, 2001; Zhang et al., 2011), and now the ABCDE model was known and applied in the studies of the flower development in plant (Pelaz et al., 2000; Pelaz et al., 2001; Theissen and Saedler, 2001; Honma and Goto, 2001; Pinyopich et al., 2003; Jack, 2004; Ditta et al., 2004; Melzer and Theissen, 2009; Varkonyi-Gasic et al., 2011). In terms of the ABCDE model, the A+E genes specific referred to the sepalswhile the C+E genes specify the carpels; the A+B+E genes to the petals; the B+C+E genes to the stamensas well; Whereas the D gene individually specify the ovules. Exception of AP2, all of the homologous flower genes are members of the plant-specific MIKC MADS-box family (Ma and dePamphilis, 2000; Kaufmann et al., 2005; Adam et al., 2007; Xu et al., 2008; Varkonyi- Gasic et al., 2011). The MADS-box genes are vital in the formation of the floral meristem, development of the floral organs and florescence regulation in plants (Ng and Yanofsky et al., 2001; Becker and Theissen, 2003; Angenent et al., 2006; Guo, 2008; van Dijk and van Ham, 2010). In recent years, many studies have focused on the facets of the cloning, functional and comparative genomics and molecular evolution of the MADS-box genes.
Ornamental plants in the Guzmania genus, which belong to the Bromeliaceae family, are native to the American tropics and subtropics as well as are sold as superior-quality potted flowering plants. There is currently a lack of information on the Guzmania MADS-box gene. In this study, we atemped to clone the MADS-box gene from a Guzmania variety, Ostara, and analyze and express the genes in different tissues in Ostaras, In order to figure out the mechanisms of gene action in the flower development of Guzmania plant, which would be helpful to understand the floral meristem formation, development of floral organs, regulation of flowering time, and the molecular regulation and control of flower shape in Guzmania varieties.
1 Results
1.1 Cloning of the full-length cDNA sequence of GoMADS-box2
A 948 bp length cDNA sequence was assembled based on four EST sequences previously obtained from a full-length EST sequence library. To further validate the sequence, we completely sequenced ppfca 0_0002_G 08.z1.ab 1, one of the above 4 EST clonesby using a primer walking approach. BLASTp analysis showed that the 985 bp length sequence was identical to the above assembled 948 bp length sequence; therefore, we considered that both of sequences were the same sequence of the gene. The 985 bp cDNA sequence contains a 675 bp ORF (open reading frame) from the start of the 75th base to the end of the 752th base that encodes 225 amino acid residuals (Figure 1), which was named as GoMADS- box2 gene and deposited in GenBank with Accession number JN936044.

1.2 Sequence analysis of the GoMADS-box2 gene
1.2.1 Blast analysis
Blast analysis of nucleotide to nucleotide showed that the full-length cDNA sequence of the GoMADS-box2 gene had a different extent of sequence identity to the MADS-box genes of Agave tequilana (81%), Alstroemeria ligtu (79%), Galeola falconeri (79%), Paphiopedilum (79%), Hypoxis villosa (79%), Tricyrtis affinis (80%), and Calycanthus floridus (79%), in which the sequence had the hightest homology to Agave tequilana (gbJF699271.1). While by comparing the deduced amino acid sequences (BLASTp) in the protein data base, the results exhibited that the GoMADS-box2 protein had higher sequence identity to the MADS-box proteins of other species that Muscari armeniacum (dbj BAE 48147.1) had 88% amino acid homology was 88%.
1.2.2 Characteristic analysis of the deduced amino acid sequence of GoMADS-box2
ProtParam analysis indicated that the theoretical molecular weight and isoelectric point of the GoMADS-box2 gene be 26.3 kD and 9.36, respectively, and there were 28 negatively charged amino acids (Asp+Glu) as well as 36 positively charged amino acids (Arg+Lys). While secondary structure analysis with the SPOMA program revealed that the protein consisted of alpha helices (56.00%), random coils (24.44%), extended strands (13.78%) and beta turns (5.78%) (Combet et al., 2000). Conservative domain analysis by using the CDART program (NCBI) figured out that there were two typical conserved domains, the MADS and K-box domains in the gene (Figure 2) (Marchler-Bauer et al., 2011; Marchler- Bauer et al., 2009; Marchler-Bauer and Bryant 2004). The structures of MADS, I, K-box and C domains were located in AA2-60, AA61-70, AA71-170 and AA171-225, respectively (Figure 1), which should be the characteristics of plant-specific MIKC-type MADS-box proteins (Zheng et al., 2004; Folter et al., 2005; Immink et al., 2010; van Dijk et al., 2010).
![]()
1.2.3 Tertiary structure of GoMADS-box2 protein
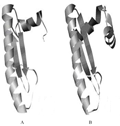
Based on the 3D structure of the 1TQE P chain, a tertiary structure model was generated by the ESyPred 3D program (Figure 3), which determined that there was 19.1% homology between the 1TQE Pchain and the GoMADS-box2 protein (Lambert et al., 2002). The 2PSN structure, which was obtained by Han et al., in 2005 (Han et al., 2005), consisted of P, Q, R, S, X, and Y chains, and the P, Q, R, S chains were annotated as "transcription factor, MADS-box" by the InterPro program.
1.2.4 Molecular phylogenetic tree of GoMADS-box2
To confirm the class that GoMADS-box2 gene belongs to, molecular evolution analysis was performed byusing the ABCDE model (Colombo et al., 1995; Pelaz et al., 2000; Xu et al., 2008). The evolutionary tree was generated by the MEGA 4.1 software with the neighbor-joining (NJ) method after a complete alignment was performed by using the Clustalx1.81 program. The results from the analysis were shown in Figure 4. Similar to AP3 (GI5805228, AAD51896.1) and PI (GI642128, D30807.1) of Arabidopsis thaliana, the GoMADS-box2 gene was assigned to B-class gene. Consequently, we thought that this gene might play an important role in the regulation and control of petal and stamen development.
Furthermore, molecular evolution analysis by using Theissen's classification method (Theissen et al., 2000) revealed that the MADS-box proteins contained AP3, GLO, AG, AGL6, SQUA, and AGL2 (Figure 5). Consistent with our expectations, different MADS- box proteins were grouped into different branches. However, the GoMADS-box2 proteins, Apetela3 (GI166607) of Arabidopsis thaliana, Def (GI110671803) and Tm 6 (GI110671805) of Petunia were clustered together in the AP3 group.
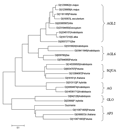

Figure 5 Molecular phylogenetic tree of GoMADS-box2 based on Theissen's classification method (Theissen et al., 2000)
1.3 Tissue expression of the GoMADS-box2 gene
The standard curves of Goactin1, GoGAPDH1, and GoMADS-box2 were made as follows: Goactin1: y= -3.218x+15.938, R2=0.990; GoGAPDH1: y=-3.314x+ 16.074, R2=0.992; GoMADS-box2: y=-3.172x+16.572, R2= 0.996. Because all of the R2 values were 0.99 above and almost close to 1.00, a strong correlation between the Ct values and logarithmic values of the template’s initial copy numbers exsited. Meanwhile, a gel electrophoresis and melting curve test also confirmed the results of the real-time quantitative PCR experiment. The Ct values of each gene were generated by three parallel quantitative PCR reactions, and the copy numbers were calculated based on the equations of their standard curve. Finally, after normalizing the copy number of the GoMADS-box2 gene by using two internal reference genes (Goactin1 and GoGAPDH1), we determined the relative expression levels of GoMADS-box2 in various tissues (Figure 6), the results showed that GoMADS-box2 was expressed in all of the tested tissues, in which the floral expression level was the highest than that of other three tissues and 10 times more than that of the scape. According to t-test analysis between scape and leaf expression levels by using the PASW Statistics software (Ver. 18), the p value reached 0.307, indicating that was much more at 0.05 level, obviously, there was no significant difference between the two tissues. Whereas the relative expression level in the bract lowered to zero.
![]()
2 Discussions
The MADS-box, a highly conserved DNA domain, was found in the mini chromosome maintenance 1 gene (MCM1) of yeast, agamous gene (AG) of Arabidopsis thaliana, deficiens gene (DEF) of Antirrhinum majus, and serum response factor gene (SRF) in humans (Theissen et al., 1996; Theissen et al., 2000; Zheng et al., 2004). The MADS-box protein exists in many animals and plants. The MADS-box genes in higher plants are composed of M, I, K, C domains with different degrees of conservation. The MADS-box domain (M) consists of 60 amino acids and is the most conserved (Purugganan et al., 1995). The K domain (keratin structure domain) is located from 65 to 70 amino acids downstream and is moderately conserved (Ma, 1991). The intervening region(I) is located at 35 amino acids between the MADS-box and K domains, and the carboxyl-terminal (C) domain, which is located at 30 amino acids downstream of the K domain, is the least conserved (Ma, 1991). The MADS and K-domains are involved in mediating protein-protein interactions (Davies et al., 1996), and the I-domain was thought to be important for interaction specificity (Krizek and Meyerowitz, 1996; Riechmann et al., 1996; van Dijk et al., 2010).
The GoMADS-box2 gene is composed of a highly conserved M domain with 59 amino acids, a moderately conserved K domain with 100 amino acids, and I domain with 10 amino acids and a C domain with 55 amino acids; this structure is characterized ed as the MIKC type MADS-box protein, which only exists in plants. With the exception of the M domain, the amino acid numbers of the K, I, and C domains were different from other reported MADS-box proteins in plants. However, an analysis of the sequence characteristics of the AP3 genes demonstrated that, SsMADS, the AP3 homolog gene in Salix suchowensis, was very similar to the GoMADS-box2 gene. For example, the amino acid numbers of the M, K, I and C domains in the SsMADS gene were 59, 100, 10 and 60, respectively. There was a single methionine residue that extended outward in front of the M domain, which is consistent with the GoMADS-box2 sequence (Chen et al., 2007). Hence, the structural characteristics of the amino acid sequence in various conserved domains in AP3-class genes differ to some extent from those of other MADS-box genes.
According to the analysis of the molecular phylogenetic tree, theGoMADS-box2 gene belongs to the B-class genes and AP3-type genes. Moreover, based on previous research, the B-class genes include Apetala3 (Ap3), Deficiens (Def), Pistillata (Pi) and Globosa (Glo).The Ap3 gene is a B-type gene, which is indirectly supported by the former results (Chen et al., 2007).
Housekeeping genes often act as internal reference genes in real-time quantitative PCR. However, according to some reports, the expression levels of some housekeeping genes may change in different conditions (Tu et al., 2007). To assure test reliability, two housekeeping genes, ACTIN (Goactin1) and GAPDH (GoGAPDH1), were used to normalize the expression levels of GoMADS-box2. According to the experimental results, the expression levels of GoMADS-box2 in the flower are greater than that the levels in the other three tissues. Because GoMADS- box2 may not only regulate floral specificity butalso is a B-type gene that controls petal and stamen development, the high amount of expression of GoMADS-box2 in flowers is consistent with our expectations.
In this paper, the full-length cDNA sequence of the GoMADS-box2 gene was cloned in Guzmania Ostara and exhibited characteristics of both AP3-type and B-class genes that control petal and stamen development. Furthermore, this gene was expressed in all of the tested tissues with the highest expression levels in the flower. The findings from this study would be valuable in understanding the molecular regulation and control of flower shape in Guzmania.
3 Materials and Methods
3.1 Cloning of GoMADS-box2
In 2008, we successfully constructed a full-length cDNA library by using the floral organs and bracts of Ostara, which consisted of 1758 high-quality sequences through 5'EST sequencing 2004 positive clones that were picked at random (Liu et al., 2009). After all of the sequences were analyzed by Blast, Four ESTs were screened in the same contig that harbored the MADS-box genes by blasting all the sequenced cDNA, including ppfca0_000788.z1.scf, ppfca0_000979.z1.scf, ppfca0_0002_B09.ab1, and ppfca0_0002_G08.z1.ab1. Full-length cDNA sequences of the MADS-box genes were then obtained via two strategies. One of the cDNA sequence of 948 bp in length was assembled by the 4 EST sequences Another cDNA sequence of 985 bp in length was generated by using the primer walking sequencing approach on the clone of ppfca 00002_G08.z1.ab, one of four EST clones.
3.2 Sequence analysis of theGoMADS-box2 gene
The characteristics of the cDNA sequence, amino acid residuals and molecular phylogenetic tree of the GoMADS-box2 gene were analyzed by using online bioinformatics tools such as BlastN, BlastP, ExPASy, ProtParam, SPOMA, CDART(NCBI), ESyPred 3D, ClustalX (1.81) and MEGA 4.1 software.
3.3 Tissue expression of GoMADS-box2
3.3.1 Primer design
Two housekeeping genes, actin and GAPDH, were served as internal references to normalize the amount of expression of the tested gene in the experiment. Both Goactin1 (GenBank accession number: HQ184438) and GoGAPDH1 (GenBank accession number: HM185058) were previously obtained from Guzmania Ostara. The primers used were as follows: Goactin1-F (forward primer): 5'-GCTTGCCTACATT GCCCT-3'; Goactin1-R (reverse primer): 5'-ATTGT TGAACCCCCGCTT-3'; GoGAPDH1-F: 5'-CAAC TGTCCTGCTCCTCTA-3'; GoGAPDH1-R: 5'-GGC AAGTCAAGTCCACAAC-3'; GoMADS-box2-F: 5'- AACCTCCGCAAAGAAATA-3'; GoMADS-box2-R: 5'-ATCCACGAAGCCATAGAC-3'.
3.3.2 Extraction and reverse transcription of total RNA
Following the TRIzol reagent’s user's manual (Dingguo, China), whole RNA was extracted from the materials of scape, leaf, bract and flower of Ostara; then the concentration, purity and integrity were analyzed by ultraviolet spectrometry as well as agarose gel electrophoresis. Further, the total RNA was reverse transcribed into cDNA following the user's manual of the First Strand cDNA Synthesis Kit (Fermentas K162, Canada).
3.3.3 Preparation of standard curves
First, the GoMADS-box2, Goactin1, and GoGAPDH1 genes were individually amplified by PCR (System 9600, Perkin Elmer), then these PCR products were diluted tenfoldafter being identified by agarose gel electrophoresis. Finally four consecutive concentra- tions were selected as the standards to prepare the standard curves.
3.3.4 Real-time quantitative PCR analysis
The 25 µL volume real-time quantitative PCR reaction included 1 µL cDNA, 0.5 µL primer-F (20 pmol/µL), 0.5µl primer-R (20 pmol/µL), 12.5 µL 2×mix, 1 µL SybrGreen I (10×concentration) and 9.5 µL ddH2O. The PCR procedures were listed in Table 1 performed in PRISM 7700 Sequence Detector ABI.
Table 1 Quantitative PCR procedures
|
3.3.5 Operation of apparatus
With the complete of the PCR preparation and procedures, the PCR amplification was performed on a 96-well plate in the Real-time Quantitative PCR System of ABI 7 700. To ensure a correlation coefficient closed to 1.00, the quality and concentrations of standard samples were adjusted.
Author’s contributions
Author’s contributions Jianxin Liu was deviser, superintendent and principal transactor; Weiyong Wang, Fuquan Shen and Huaqiao Ding provided test materials and analyzed data. FeiZhang and Xiaojing Liu participated in experimental design, test result analysis and writing of the preliminary draft; All authors had read and approved the final version.
Acknowledgements
This study was jointly supported by the Great Science and Technology Special Project of the Zhejiang Province (No.2009C12095) and the Technological Innovation Ability Promoting Project and Initiation Project of Doctoral Scientific Research in the Zhejiang Academy of Agricultural Sciences.
References
http://dx.doi.org/10.1093/jxb/erl263
http://dx.doi.org/10.1016/S1055-7903(03)00207-0
Coen E.S., and Meyerowitz E.M., 1991, The war of the whorls: Genetic interaction controlling flower development, Nature, 353: 31-37
http://dx.doi.org/10.1038/353031a0
Colombo L., Franken J., Koetje E., van Went J., Dons H.J., Angenent G.C., and van Tunen A.J., 1995, The petunia MADS box gene FBP11 determines ovule identity, Plant Cell, 7: 1859-1868
Combet C., Blanchet C., Geourjon C., and Deléage G., 2000, NPS@: Network protein sequence analysis, Tibs. 25: 147-150
http://dx.doi.org/10.1016/S0968-0004(99)01540-6
Davies B., EgeaCortines M., Silva E.D., Saedler H., and Sommer H., 1996, Multiple interactions amongst floral homeotic MADS box proteins, Embo Journal, 15: 4330-4343
Ditta G., Pinyopich A., Robles P., Pelaz S., and Yanofsky M.F., 2004, The SEP4 gene of Arabidopsis thaliana functions in floral organ and meristem identity, CurrBiol, 14(21): 1935-1940
http://dx.doi.org/10.1016/j.cub.2004.10.028
http://dx.doi.org/10.1105/tpc.105.031831
http://dx.doi.org/10.1016/j.jmb.2004.10.033
http://dx.doi.org/10.1038/35054083
http://dx.doi.org/10.1016/j.semcdb.2009.10.004
http://dx.doi.org/10.1105/tpc.017038
Kaufmann K., Melzer R., and Theißen G., 2005, MIKC-type MADS-domain proteins: structural modularity, protein interactions and network evolution in land plants, Gene, 347(2): 183-198
http://dx.doi.org/10.1016/j.gene.2004.12.014
Krizek B.A., and Meyerowitz E.M., 1996, Mapping the protein regions responsible for the functional specificities of the Arabidopsis MADS domain organ-identity proteins, Proc. Natl. Acad. Sci., USA, 93: 4063-4070
http://dx.doi.org/10.1073/pnas.93.9.4063
Lambert C., Leonard N., De Bolle X., and Depiereux E., 2002, ESyPred3D: Prediction of proteins 3D structures, Bioinformatics, 18(9): 1250-1256
http://dx.doi.org/10.1093/bioinformatics/18.9.1250
http://dx.doi.org/10.1101/gad.5.3.484
Ma H., and dePamphilis C., 2000, The ABCs of floral evolution, Cell, 101(1): 5-8
http://dx.doi.org/10.1016/S0092-8674(00)80618-2
Marchler-Bauer A., Anderson J., Chitsaz F., Derbyshire M.K., DeWeese-Scott C., Fong J.H., Geer L.Y., Geer R.C., Gonzales N.R., Gwadz M., He S., Hurwitz D.I., Jackson J.D., Ke Z., Lanczycki C.J., Liebert C.A., Liu C., Lu F., Lu S.,
Marchler G.H., Mullokandov M., Song J.S., Tasneem A., Thanki N., Yamashita R.A., Zhang D., Zhang N., and Bryant S.H., 2009, CDD: specific functional annotation with the Conserved Domain Database, Nucleic Acids Res., 37(D): 205-210
http://dx.doi.org/10.1093/nar/gkn845
Marchler-Bauer A., and Bryant S.H., 2004, CD-Search: protein domain annotations on the fly, Nucleic Acids Res., 32(W): 327-331
http://dx.doi.org/10.1093/nar/gkq1189
Melzer R., and Theissen G., 2009, Reconstitution of 'floral quartets' in vitro involving class B and class E floral homeotic proteins, Nucleic Acids Research, 37(8): 2723-2736
http://dx.doi.org/10.1093/nar/gkp129
Ng M., and Yanofsky M.F., 2001, Function and evolution of the plant MADS-box gene family, Nat. Rev. Genet., 2(3): 186-195
http://dx.doi.org/10.1038/35056041
Pelaz S., Ditta G. S., Baumann E., Wisman E., and Yanofsky M.F., 2000, B and C floral organ identity functions require SEPALLATA MADS-box genes, Nature, 405(6783): 200-203
http://dx.doi.org/10.1038/35012103
Pelaz S., Gustafson-Brown C., Kohalmi S.E., Crosby W.L., and Yanofsky M.F., 2001, APETALA1 and SEPALLATA3 interact to promote flower development, Plant J., 26(4): 385-394
http://dx.doi.org/10.1046/j.1365-313X.2001.2641042.x
Pinyopich A., Ditta G.S., Savidge B., Liljegren S.J., Baumann E., Wisman E., and Yanofsky M.F., 2003, Assessing the redundancy of MADS-box genes during carpel and ovule development, Nature, 424(6944): 85-88
Purugganan M.D., Rounsley S.D., Schmidt R.J., and Yanofsky M.F., 1995, Molecular evolution of flower development: diversification of the plant MADS-box regulatory gene family, Genetics, 140: 345-356
Riechmann J.L., Krizek B.A., and Meyerowitz E.M., 1996, Dimerization specificity of Arabidopsis MADS domain homeotic proteins APETALA1, APETALA3, PISTILLATA, and AGAMOUS, Proc. Natl. Acad. Sci., USA, 93: 4793-4798
http://dx.doi.org/10.1073/pnas.93.10.4793
Theissen G., Becker A., Rosa A.D., Kanno A., Kim J.T., Münster T., Winter K.U., and Saedler H., 2000, A short history of MADS-box genes in plant, Plant Molecular Biology., 42: 115-149
http://dx.doi.org/10.1023/A:1006332105728
Theissen G., Kim J., and Saedler H., 1996, Classification and phylogeny of the MADS-box multigene family suggest defined roles of MADS-box gene subfamilies in the morphological evolution of eukaryotes, Journal of Molecular Evolution, 43: 484-516
http://dx.doi.org/10.1007/BF02337521
Theissen G., and Saedler H., 2001, Plant biology: Floral quartets Nature, 409(6819): 469-471
http://dx.doi.org/10.1038/35054172
Tu L.L., Zhang X.L., Liu D.Q., Jin S.X., Cao J.L., Zhu L.F., Deng F.L., Tan J.F., and Zhang C.B., 2007, Suitable internal control genes for qRT-PCR normalization in cotton fiber development and somatic embryo genesis. Chin Sci. Bull., 52: 3110-3117
http://dx.doi.org/10.1007/s11434-007-0461-0
van Dijk A.D.J., Morabito G., Fiers M., van Ham R.C.H.J., Angenent G.C., and Immink R.G.H., 2010, Sequence motifs in mads transcription factors responsible for specificity and diversification of protein-protein interaction, PLoS
Comput Biol., 6(11): e1001017
http://dx.doi.org/10.1371/journal.pcbi.1001017
van Dijk A.D.J., and van Ham R.C.H.J., 2010, Conserved and variable correlated mutations in the plant MADS protein network, BMC Genomics., 11: 607
http://dx.doi.org/10.1186/1471-2164-11-607
http://dx.doi.org/10.1186/1471-2229-11-72
http://dx.doi.org/10.1016/0092-8674(94)90291-7
Xu Q.J., Guan L.F., Wu X.N., Sun L., Tan W.B., Nie Y.Z., and Li Y.H., 2008, Cloning and expression analysis of MADS-box genes from Lisianthus (Eustomagrandiflorum), Chinese Bulletin of Botany, 25(4): 415-429 (in Chinese)
Zhang Y., Wang X., Zhang W., Yu F., Tian J., Li D., and Guo A., 2011, Functional analysis of the two brassica AP3 genes involved in apetalous and stamen carpelloid phenotypes, PLOS ONE, 6(6): e20930
http://dx.doi.org/10.1371/journal.pone.0020930
. PDF(1648KB)
. FPDF
. HTML
. Online fPDF
Associated material
. Readers' comments
Other articles by authors
. Jianxin Liu
. Weiyong Wang
. Fuquan Shen
. Huaqiao Ding
. Fei Zhang
. Xiaojing Liu
Related articles
. Guzmania Ruiz & Pav
. GoMADS-box2
. real-time quantitative PCR
Tools
. Email to a friend
. Post a comment



